Did you know that 65% of the Human population has a reduced ability to digest lactose after infancy? Inversely, only about 5% of people in Northern European descent are lactose intolerant. But when you look at the Chinese population, over 90% are lactose intolerant. How can this be?
It’s because of the LCT/MCM6 gene — LCT is one of my favorite genes! This gene is so interesting because it contains an important genetic polymorphism (rs4988235) that are highly variable in different populations. This genetic polymorphism causes lactase persistence.
What Is Lactase Persistence?
Lactase-persistence is the continued function of the lactase enzyme in adulthood. If you are lactase persistent, you are (usually) able to drink milk as an adult.
Since it’s more common to be lactose intolerant in most populations, the genetic condition for being able to tolerate lactose into adulthood is referred to as lactase persistence. When looking at genetic population frequencies, the allele frequency represents persistence of the lactase enzyme as opposed to a decrease in lactase function (intolerance). A higher allele frequency means more persistence of the lactase enzyme in a population and lower frequency means less persistence of the lactase enzyme in a population.
Lactase Persistence Alleles By Population
| Population | Allele Frequency |
|---|---|
| African | 12.3% |
| European (Finnish) | 57.1% |
| South Asian | 0% |
| Other | 47.6% |
| East Asian | 0.06% |
| Ashkenazi Jewish | 10% |
| European (non-Finnish) | 60.1% |
| Latino | 20.3% |
| Male | 40.59% |
| Female | 42.8% |
| Total | 41.6% |
Data source: gnomAD v2.1.1
If you were to use this single genetic polymorphism as an estimate of lactose intolerance, you would assume that lactose intolerance is the least for non-Finnish European and the most for South and East Asian. There is another genetic polymorphism (rs182549) that is part of the haplotype, but since the allele frequencies are extremely similar, I am not going to include a table in this article.
This table shows that 12.3% of Africans have one or more alleles for lactase persistence, 57.1% of Finnish Europeans have alleles for lactase persistence, 0% of South Asians have alleles for lactase persistence, 0.06% of East Asians have alleles for lactase persistence, 10% of Ashkenazi Jews have alleles for lactase persistence, 60.1% of non-Finnish Europeans have alleles for lactase persistence and 20.3% of Latinos have alleles for lactase persistence. In other words, Asians are the most likely population to be lactose intolerant because of decreased activity of the lactase enzyme and non-Finnish Europeans are the least likely to be lactose intolerance from persistence of the lactase enzyme.
Estimated lactose intolerance genetics by ehtnicity from most intolerance to to least intolerance (data from gnomAD):
- South Asian
- East Asian
- Ashkenazi Jewish
- African
- Latino
- European (Finnish)
- European (non-Finish)
Comparing Lactose Intolerance Genetic Polymorphisms to a Worldwide Prevalence Map of Lactose Intolerance
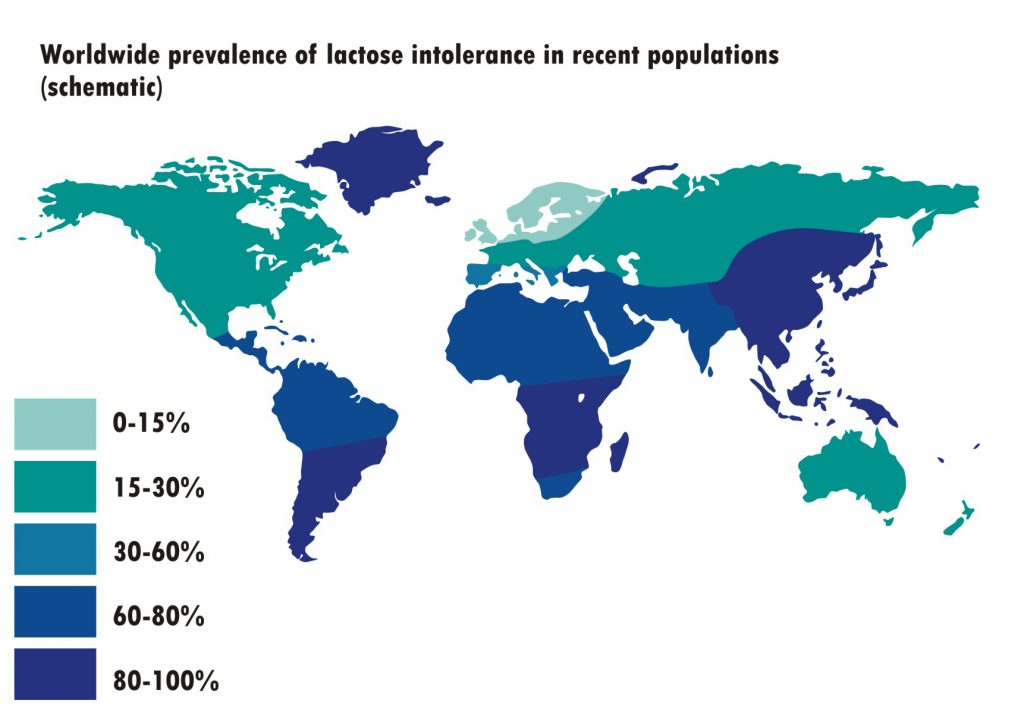
The table does not represent all populations, but for the populations it represents, the data correlates with this map well. As the map shows, the United States and Canada has roughly 15-30% lactose intolerance, East Asia have 80-100% lactose intolerance, South Asia has 60-80% lactose intolerance, African has between 30-80% lactose intolerance depending on region, and Northern Europe has the lowest amount of lactose intolerance at 0-15%. Interestingly, while Latinos are predicted to be more tolerant than Africans by genetics, 60-100% of the Latino population is predicted to be lactose intolerant. However the northern areas of Mexico are predicted to have a similar amount of lactose intolerance as the United States so perhaps the Latino allele frequencies are the result of sampling bias.
With this map, one can assume that Australia and Russia’s prevalence of the polymorphisms are probably pretty similar to United States and Europe.
Here’s a table of lactose intolerance by country:
| Country | Prevalence |
|---|---|
| Afghanistan | 82% |
| Algeria | 62% |
| Angola | 94% |
| Armenia | 98% |
| Australia | 44% |
| Austria | 22% |
| Azerbaijan | 96% |
| Belgium | 15% |
| Botswana | 88% |
| Brazil | 60% |
| Cambodia | 68% |
| Cameroon | 89% |
| Canada | 59% |
| Chile | 56% |
| China | 85% |
| Colombia | 80% |
| Cyprus | 16% |
| Czech Republic | 81% |
| Democratic Republic of the Congo | 95% |
| Denmark | 4% |
| Egypt | 68% |
| Estonia | 28% |
| Ethiopia | 77% |
| Finland | 19% |
| France | 36% |
| Gabon | 93% |
| Germany | 16% |
| Ghana | 100% |
| Greece | 55% |
| Hungary | 39% |
| India | 61% |
| Iran | 88% |
| Iraq | 93% |
| Ireland | 4% |
| Israel | 89% |
| Italy | 72% |
| Japan | 73% |
| Jordan | 56% |
| Kazakhstan | 75% |
| Kenya | 39% |
| Kuwait | 56% |
| Lebanon | 78% |
| Malawi | 100% |
| Malaysia | 87% |
| Mexico | 48% |
| Mongolia | 88% |
| Morocco | 73% |
| Mozambique | 95% |
| Myanmar | 92% |
| Namibia | 93% |
| Netherlands | 12% |
| New Zealand | 10% |
| Niger | 13% |
| Nigeria | 87% |
| Norway | 12% |
| Oman | 96% |
| Pakistan | 58% |
| Papua New Guinea | 91% |
| Poland | 43% |
| Portugal | 40% |
| Republic of Congo | 93% |
| Russia | 61% |
| Rwanda | 49% |
| Saudi Arabia | 28% |
| Senegal | 79% |
| Solomon Islands | 99% |
| Somalia | 94% |
| South Africa | 81% |
| South Korea | 100% |
| Spain | 29% |
| Sri Lanka | 73% |
| Sudan | 55% |
| Sweden | 7% |
| Syria | 95% |
| Taiwan | 88% |
| Tanzania | 45% |
| Thailand | 84% |
| Tunisia | 84% |
| Turkey | 69% |
| Uganda | 87% |
| Ukraine | 61% |
| United Kingdom | 8% |
| United States | 36% |
| Uruguay | 65% |
| Uzbekistan | 92% |
| Vietnam | 98% |
| Western Sahara | 53% |
| Yemen | 100% |
| Zambia | 98% |
Data Source: milk.procon.org
Even if there isn’t great genetic population data for every country, I think it’s safe to assume that an increase in prevalence of lactose intolerance means the population has less of the genetic polymorphisms that result in lactase persistence. Inversely, a decrease in prevalence of lactose intolerance probably means that there is a higher degree of genetic polymorphisms for lactase persistence.
Are Genetics the Cause of Lactose Intolerance?
Are genetic polymorphisms a perfect predictor of lactase intolerance or lactase-persistence? The NIH’s Genetics Home Reference (GHR) states, “Most people with lactase nonpersistence retain some lactase activity and can include varying amounts of lactose in their diets without experiencing symptoms.” While some with lactase nonpersistence may not experience symptoms of lactose intolerance with dairy consumption, these two polymorphisms are still highly predictive of lactose intolerance. There can be primary and secondary causes of lactose intolerance as well. According to Wikipedia, secondary causes of lactose intolerance are “acute gastroenteritis, coeliac disease, Crohn’s disease, ulcerative colitis, chemotherapy, intestinal parasites (such as giardia), or other environmental causes.”
If you expected these polymorphisms to be 100% predictive, you would expect some Asian countries to be 100% lactose intolerant. While 90% of China’s population is lactose intolerant, it’s not 100% like some people would be led to believe by these genetic polymorphisms. However, if you have these genetics and symptoms of lactose intolerance, you likely are deficient in the enzyme lactase to varying degrees.
If you have a 23andMe kit, you can review your lactose intolerance here or browse your 23andme raw data to look directly at the polymorphisms rs4988235 and rs182549. Are you lactose intolerant and does your genetics reflect it? Contrary, are you lactose tolerant but have the genetics for lactose intolerance?
The data is unbelievably skewed
Azerbaijan is shown as having 96% intolerance rate which absolutely ridiculous, milk, cheese, butter is #1 food consumed there
First not all dairy products have lactose content. It’s only lactose which is the sugar found in milk. It’s a disaccharide containing glucose and galactose.
Just because a population consumes dairy products does not mean they are not intolerant.
FYI cheese and butter have very little and no lactose respectively.
BTW milk cheese butter are 3 separate foods so sort of impossible for them to all be the number 1 food consumed.
Might want to actually have basic info of lactose content before making claims about data accuracy. Sort of hard to “skew” DNA population data. It is what it is.
BTW even combining all common milk products together its still not even close to the number one food consumed in Azerbaijan. Just as an example only (all numbers wre thousand so add 000) 790 tons of milk was produced there. It has been steadily dropping since the 1980s when is was around 150,000 ton As a comparison 125,000 t of chicken, 110,000 t of eggs for Azerbaijan.
The number 1 food consumed in Azerbaijan is rice by far. You would think any native would know that. Besides meat, rice, eggs, bread all easily surpass milk product consumption and again let’s not forget we are speaking specifically about the data on lactose so only soft cheese little and actual milk and yogurt for direct products.
I am homozyous to both these rs #’s by testing on 23andMe and told I have genetic lactose intolerance since birth by my doctor. Upper endoscopy was 2.2 with lactase (normal is anything above 15 with lab). Also, I am close to one quarter Ashkenazi Jewish DNA. To be comfortable I find I must be 100% dairy free.
Koreans are 100% on the table, but most of them drink milk well. Why? I didn’t even know lactose intolerance existed 5 years ago. Because everyone drinks milk in Korea.
this makes me wonder if there are other genes that certain populations have that have similar encoding of the LCT gene
Im heterozygous but I still have digestion issues with milk and cheeses. I’ve had it since I was 7 years old (gas, bloating, diarrhea). I am also Jewish/middle eastern.
What’s your origin? Ashkenazi?
heterozygotism means you have less tolerance compared to homozygotic dominant, considering there is literally one less gene fulfilling a certain task. This is true for many things.